Concept 3.4: Regulation of Metabolism Occurs by Regulation of Enzymes
The enzyme-catalyzed reactions we have been discussing often operate within metabolic pathways in which the product of one reaction is a substrate for the next. For example, the pathway for the catabolism of sucrose begins with sucrase and ends many reactions later with the production of CO2 and H2O. Energy is released along the way. Each step of this catabolic pathway is catalyzed by a specific enzyme:
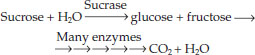
Other enzymes participate in anabolic pathways, which produce relatively complex molecules from simpler ones. A typical cell contains hundreds of enzymes that participate in many interconnecting metabolic pathways, forming a metabolic system (FIGURE 3.17). Consider a single molecule in the midst of this map:
- There may be two or more enzyme-catalyzed reactions affecting it: either making it or metabolizing it.
- Other pathways affect the concentrations of the substrates and products of these reactions.
- Each enzyme-catalyzed reaction has its own rate, depending on these concentrations.
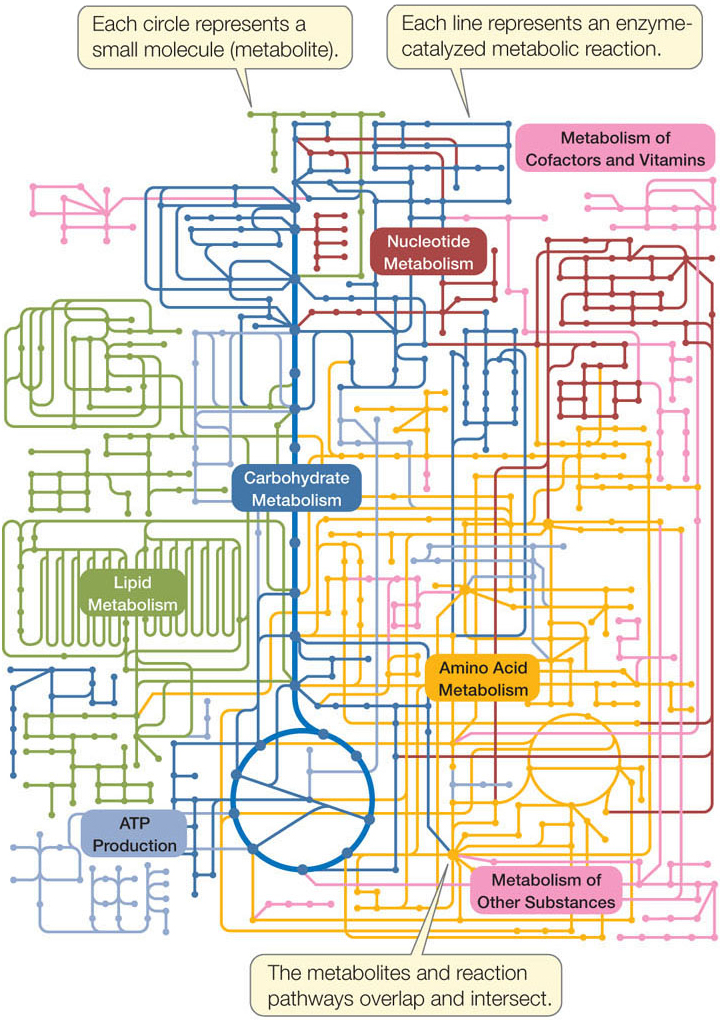
Clearly, every component of this complex system is affected by every other component, making it difficult to predict what would happen if one or more components were altered. In the new field of systems biology, scientists describe mathematically the components of metabolic systems—the concentrations of all the reactants and the rates of the reactions—and use computer algorithms to make predictions about what would happen if a component of the system were altered (see Concept 1.2).
Cells need to maintain stable internal conditions, including constant levels of certain metabolites. In addition, cells need to regulate their metabolic pathways to respond to changes, either within the organism or in its environment. One way a cell can regulate its metabolism is to control the amount of an enzyme. For example, the product of a metabolic pathway may be available from the cell’s environment in adequate amounts. In this case, it would be energetically wasteful for the cell to continue making large proteins (as most enzymes are) that it doesn’t need. For this reason, cells often have the ability to turn off the synthesis of certain enzymes.
LINK
The amount of an enzyme is controlled by regulating the expression of gene(s), a topic covered in Chapter 11
54
The consequences of too little enzyme can be significant. For example, in humans sucrase is important in digestion. In rare cases, infants are born with a congenital sucrase deficiency and the pathway that begins with sucrose is essentially blocked. If these infants ingest foods containing sucrose, the sucrose accumulates rather than being catabolized, and the infant gets diarrhea and stomach cramps. In some cases this leads to slower growth. This deficiency can be treated by limiting sucrose consumption or taking tablets that contain sucrase at every meal.
Cells can also maintain stable internal conditions by regulating the activity of enzymes. An enzyme protein may be present continuously, but it may be active or inactive depending on the needs of the cell. Synthesizing and breaking down enzymes takes time, whereas regulating enzyme activity allows cells to fine-tune metabolism relatively quickly in response to changes in the environment. In this section, we will describe how enzyme regulation occurs.
Enzymes can be regulated by inhibitors
Various chemical inhibitors can bind to enzymes, slowing down the rates of the reactions they catalyze. Some inhibitors occur naturally in cells; others can be made in laboratories. Naturally occurring inhibitors regulate metabolism; artificial ones (such as the improved version of salicylic acid described in the opening story) can be used to treat disease, kill pests, or study how enzymes work. In some cases the inhibitor binds the enzyme irreversibly, and the enzyme becomes permanently inactivated. In other cases the inhibitor has reversible effects; it can separate from the enzyme, allowing the enzyme to function fully as before.
Irreversible Inhibition
If an inhibitor covalently binds to an amino acid side chain at the active site of an enzyme, the enzyme is permanently inactivated because it cannot interact with its substrate. An example of an irreversible inhibitor is DIPF (diisopropyl phosphorofluoridate), which irreversibly inhibits acetylcholinesterase, an important enzyme that functions in the nervous system. DIPF does so by reacting with a hydroxyl group on a serine in the active site (FIGURE 3.18). The widely used insecticide malathion is a derivative of DIPF that inhibits only insect acetylcholinesterase, not the mammalian enzyme. The irreversible inhibition of enzymes is of practical use to humans, but this form of regulation is not common in the cell, because the enzyme is permanently inactivated and cannot be recycled. Instead, cells use reversible inhibition.
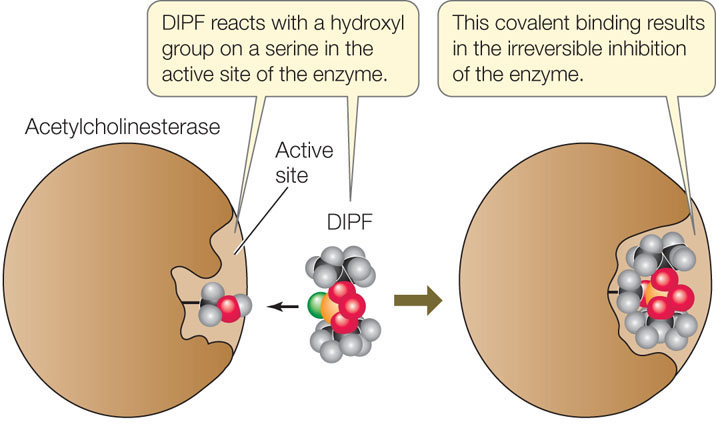
Reversible Inhibition
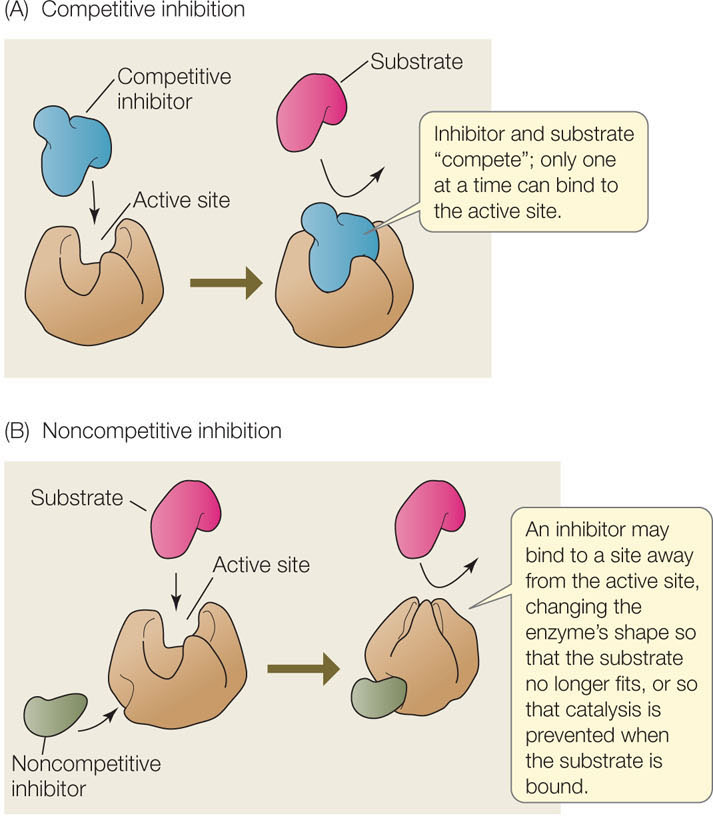
In some cases, an inhibitor is similar enough to a particular enzyme’s natural substrate that it can bind noncovalently to the active site, yet different enough that no chemical reaction occurs. This is analogous to a key that inserts into a lock but does not turn it. When such a molecule is bound to the enzyme, the natural substrate cannot enter the active site and the enzyme is unable to function. Such a molecule is called a competitive inhibitor because it competes with the natural substrate for the active site (FIGURE 3.19A). Many drugs are competitive inhibitors of enzyme targets. For example, methotrexate is a drug designed with a structure similar to the metabolite dihydrofolate. The latter is converted by an enzyme to a substance essential to cell division. Acting as a competitive inhibitor of the enzyme, methotrexate blocks cell division and is used in cancer therapy. Competitive inhibition is reversible. When the concentration of the competitive inhibitor is reduced, the active site is less likely to be occupied by the inhibitor, and the enzyme regains activity.

Go to ANIMATED TUTORIAL 3.2 Enzyme Catalysis
PoL2e.com/at3.2
A noncompetitive inhibitor binds to an enzyme at a site distinct from the active site. This binding causes a change in the shape (the conformation) of the enzyme, altering its activity (FIGURE 3.19B). The active site may no longer bind the substrate, or if it does, the rate of product formation may be reduced. Like competitive inhibitors, noncompetitive inhibitors can become unbound, so their effects are reversible.
An allosteric enzyme is regulated by changes in its shape
Noncompetitive inhibition is an example of allostery (allo, “different”; stereos, “shape”). Allosteric regulation occurs when a non-substrate molecule binds or modifies a site other than the active site of an enzyme. The site bound by the non-substrate molecule is called the allosteric site. This binding induces the enzyme to change its conformation, altering the chemical attraction (affinity) of the active site for the substrate. As a result, the rate of the reaction is changed.
An allosteric site may be modified by either noncovalent or covalent binding:
- Noncovalent binding: A regulatory molecule may bind noncovalently to an allosteric site, causing the enzyme to change shape. This noncovalent binding is reversible, and may result in the inactivation of an enzyme (see Figure 3.19B) or the activation of a formerly inactive enzyme (FIGURE 3.20A).
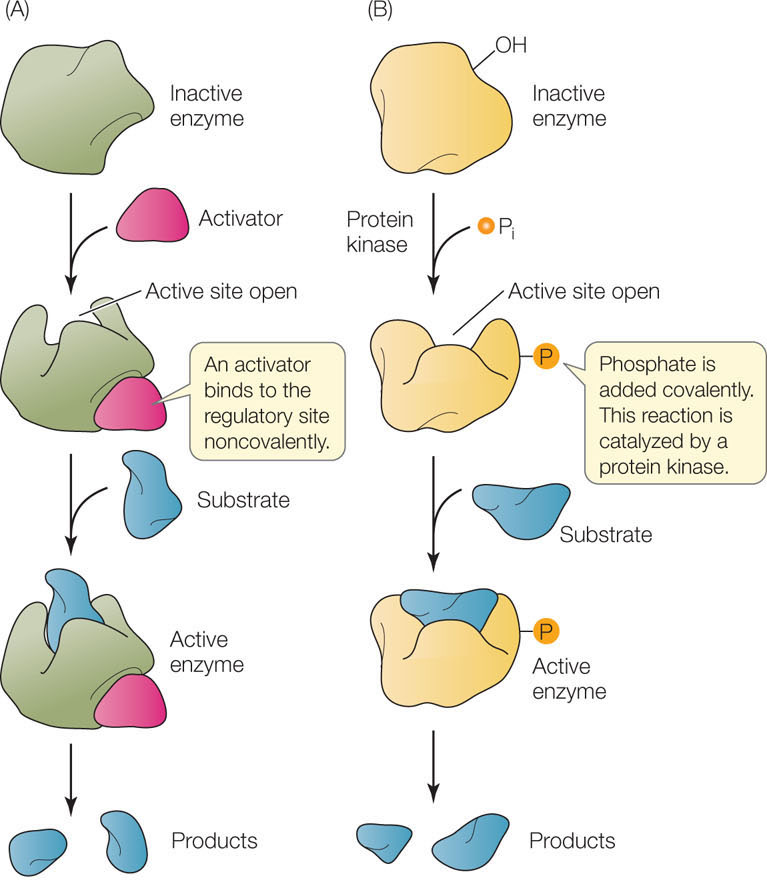 Figure 3.20: Allosteric Regulation of Enzyme Activity (A) Noncovalent binding of a regulator (in this case an activator) can cause an enzyme to change shape and expose an active site. B) Enzymes can also be activated by covalent modification, in this case phosphorylation. Note that allosteric regulation can be negative as well, with the active site becoming hidden.
Figure 3.20: Allosteric Regulation of Enzyme Activity (A) Noncovalent binding of a regulator (in this case an activator) can cause an enzyme to change shape and expose an active site. B) Enzymes can also be activated by covalent modification, in this case phosphorylation. Note that allosteric regulation can be negative as well, with the active site becoming hidden.
Go to ANIMATED TUTORIAL 3.3 Allosteric Regulation of Enzymes
PoL2e.com/at3.3
- Covalent binding: Some allosteric sites can be modified by the covalent binding of a molecule or chemical group. For example, an amino acid residue can be covalently modified by the addition of a phosphate group, in a process called phosphorylation (FIGURE 3.20B). If this occurs in a hydrophobic region of the enzyme, it makes that region hydrophilic, because phosphate carries a negative charge. The protein twists, and this can expose or hide the active site. Protein phosphorylation is an extremely important mechanism by which cells regulate many different enzymes and other proteins. It is a reversible process: a class of enzymes called protein kinases catalyze the addition of phosphate groups to proteins, whereas protein phosphatases remove phosphate groups from proteins. Humans have hundreds of different protein kinases and phosphatases. We will return to the exact functions of these proteins many times in this book.
55
LINK
Protein kinases are of particular importance in intracellular signaling pathways (see Concepts 5.5 and 5.6 and in the control of cell reproduction (see Concept 7.3)
Some metabolic pathways can be controlled by feedback inhibition
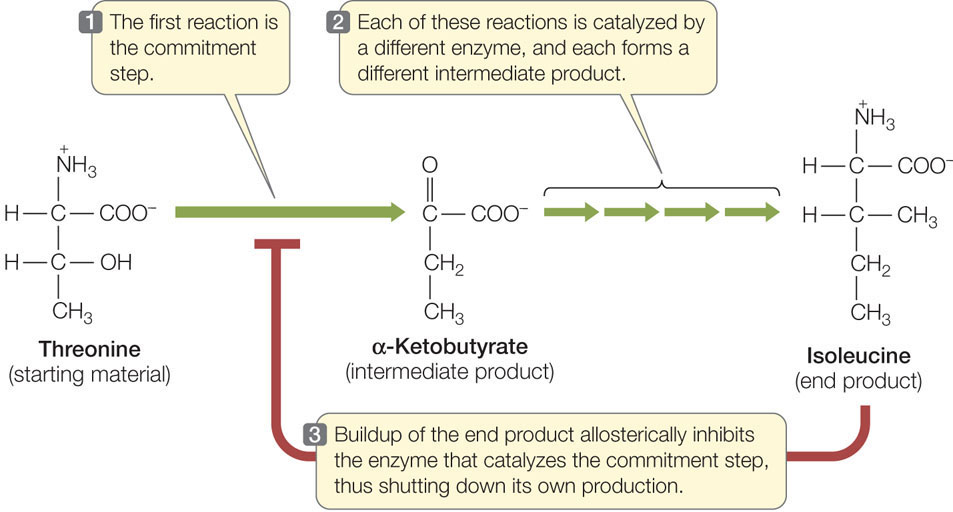
A metabolic pathway typically involves a starting material, various intermediate products, and an end product that is used for some purpose by the cell. In each pathway there are a number of reactions, each forming an intermediate product and each catalyzed by a different enzyme. In many pathways the first step is the commitment step, meaning that once this enzyme-catalyzed reaction occurs, the “ball is rolling,” and the other reactions happen in sequence, leading to the end product. But as we pointed out earlier, it is energetically wasteful for the cell to make something it does not need.
One way to regulate a metabolic pathway is by having the final product inhibit the enzyme that catalyzes the commitment step (FIGURE 3.21). When the end product is present at a high concentration, some of it binds to a site on the commitment step enzyme, thereby causing it to become inactive. The end product may bind to the active site on the enzyme (as a competitive inhibitor) or an allosteric site (as a noncompetitive inhibitor). This mechanism is known as feedback inhibition or end-product inhibition. We will describe many other examples of such inhibition in later chapters.
56
Enzymes are affected by their environment
As we have seen, the specificity and activity of an enzyme depend on its three-dimensional structure, and this in turn depends on weak forces such as hydrogen bonds (see Figure 3.7). In living systems, two environmental factors can change protein structure and thereby enzyme activity.
pH
We introduced the concept of acids and bases when we discussed amino acids. Some amino acids have side chains that are acidic or basic (see Table 3.2). That is, they either generate H+ and become anions, or attract H+ and become cations. These reactions are often reversible. For example:
Glutamic acid—COOH ⇌ glutamic acid—COO− + H+
The ionic form of this amino acid (right) is far more hydrophilic than the nonionized form (left).
From your studies of chemistry, you may recall the law of mass action or Le Chatelier’s principle. In this case the law implies that the higher the H+ concentration in the solution, the more the reaction will be driven to the left (forming more of the nonionized form of glutamic acid). Therefore changes in the H+ concentration can alter how hydrophobic some regions of a protein are and thus affect its shape. To generalize, protein tertiary structure, and therefore enzyme activity, is very sensitive to the concentration of H+ in the aqueous environment. You may also recall that H+ concentration is measured by pH (the negative logarithm of the H+ concentration).
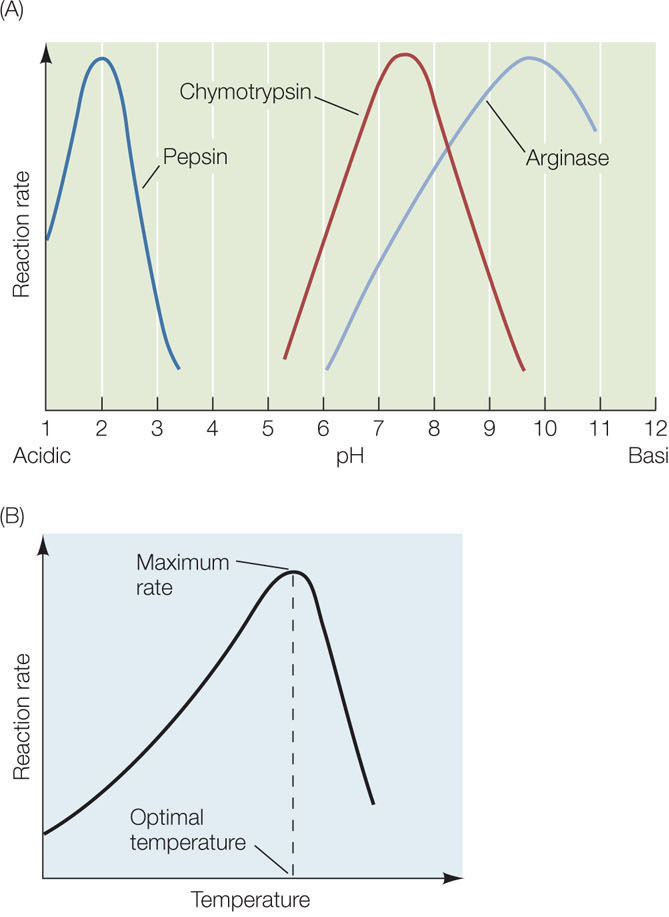
Although the water inside cells is generally at a neutral pH of 7, this can change, and different biological environments have different pH values. Each enzyme has a tertiary structure and amino acid sequence that make it optimally active at a particular pH. Its activity decreases as the solution is made more acidic or more basic than this ideal (optimal) pH (FIGURE 3.22A). As an example, consider the human digestive system (see Concept 30.4). The pH inside the human stomach is highly acidic, about pH 1.5. Pepsin, an enzyme that is active in the stomach, has a pH optimum near 2. Many enzymes that hydrolyze macromolecules in the intestine, such as proteases, have pH optima in the neutral range. So when food enters the small intestine, a buffer (bicarbonate) is secreted into the intestine to raise the pH to 6.5. This allows the hydrolytic enzymes to be active and digest the food.
APPLY THE CONCEPT: Regulation of metabolism occurs by regulation of enzymes
The concept of enzymes as biological catalysts has many applications. In a pile of clothes in your garage, you notice there are bacteria growing on some socks made of this synthetic polymer:
[—CO—(CH2)4—CO—NH—(CH2)4—NH—]n
You make a protein extract from the bacteria and isolate what you think is an enzyme that can cleave the monomers from the polymer. You also synthesize the dipeptide glycine-glycine (see Table 3.2) to test as a possible inhibitor of the enzyme. The table shows the results from several of your experiments.
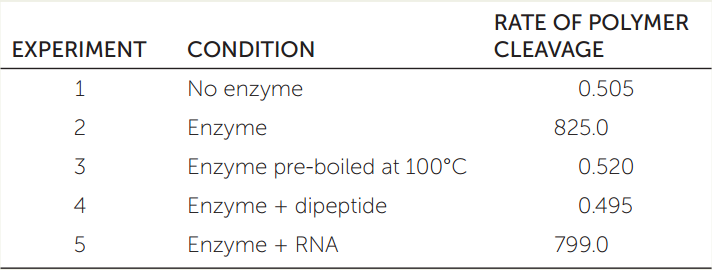

- Explain the results of each experiment.
- How might the dipeptide work? How would you test your hypothesis?
Temperature
In general, warming increases the rate of a chemical reaction because a greater proportion of the reactant molecules have enough kinetic energy to provide the activation energy for the reaction. Enzyme-catalyzed reactions are no different (FIGURE 3.22B). However, temperatures that are too high inactivate enzymes, because at high temperatures the polypetides vibrate and twist so rapidly that some of their noncovalent bonds break. When an enzyme’s tertiary structure is changed by heat, the enzyme can no longer function. Some enzymes denature at temperatures only slightly above that of the human body, but a few are stable even at the boiling point (or freezing point) of water. All enzymes have an optimal temperature for activity.
57
Individual organisms adapt to changes in the environment in many ways, one of which is based on groups of enzymes called isozymes, which catalyze the same reaction but have different chemical compositions and physical properties. Different isozymes within a given group may have different optimal temperatures. The rainbow trout, for example, has several isozymes of the enzyme acetylcholinesterase. If a rainbow trout is transferred from warm water to near-freezing water (2°C), the fish produces a different isozyme of acetylcholinesterase. The new isozyme has a lower optimal temperature, allowing the fish’s nervous system to perform normally in the colder water.
In general, enzymes adapted to warm temperatures do not denature at those temperatures because their tertiary structures are held together largely by covalent bonds such as disulfide bridges, instead of the more heat-sensitive weak chemical interactions.
CHECKpoint CONCEPT 3.4
- Explain and give examples of irreversible and reversible enzyme inhibitors.
- The amino acid glutamic acid (see Table 3.2) is at the active site of an enzyme. Normally the enzyme is active at pH 7. At pH 4 (higher concentration of H+), the enzyme is inactive. Explain these observations.
- An enzyme is subject to allosteric regulation. How would you design an inhibitor of the enzyme that was competitive? Noncompetitive? Irreversible?
- Some organisms thrive at pH 2; other organisms thrive at a temperature of 65°C. Yet mammals cannot tolerate either environment in their tissues. Explain.
Question 3.2
How does an understanding of proteins and enzymes help explain how aspirin works?
ANSWER The mechanism by which aspirin works exemplifies many of the concepts introduced in this chapter. Robert Vane showed that aspirin binds to a protein with a specific three-dimensional structure (Concept 3.2). This protein is cyclooxygenase, an enzyme (Concept 3.3) that catalyzes the commitment step in a metabolic pathway (Concept 3.4). Aspirin acts as an irreversible inhibitor of cyclooxygenase (Concept 3.4). Follow the description below carefully, as it illustrates these important concepts.
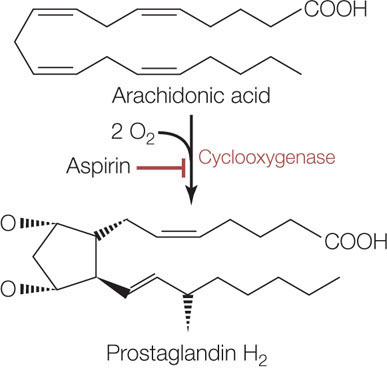
Cyclooxygenase catalyzes the conversion of a fatty acid with 20 carbon atoms, arachidonic acid, to a structure with a ring (thus the “cyclo” in the name of the enzyme). O2 is a substrate (thus the “oxygen”; FIGURE 3.23). The product of this reaction (prostaglandin H2) is the starting material for biochemical pathways that produce two types of molecules:
- prostaglandins, which are involved in inflammation and pain, and
- thromboxanes, which stimulate blood clotting and constriction of blood vessels.
58
Aspirin binds and reacts with a serine residue within the active site of cyclooxygenase. As a result of this binding, an acetyl group is transferred to the exposed hydroxyl group of the serine residue (FIGURE 3.24):
(Cyclooxygenase)-serine-OH → (cyclooxygenase)-serine-O-CH2-CH3
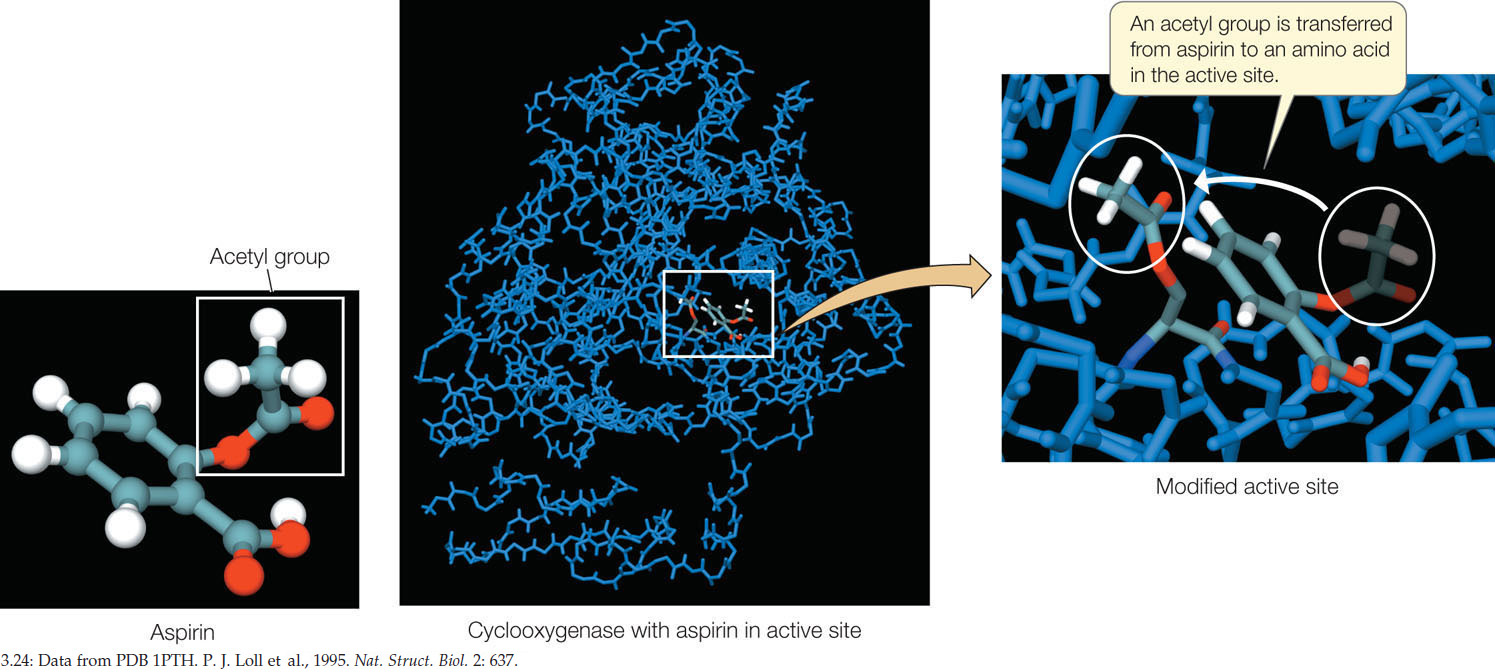
This covalent modification changes the exposed, polar serine to a less polar molecule, and it becomes slightly more hydrophobic. The conformation of the active site changes and becomes inaccessible to the substrate, arachidonic acid. The enzyme is inhibited, and the pathways leading to prostaglandins and thromboxanes are shut down. Less pain, inflammation, and blood clotting are the result. Small wonder that aspirin is taken as a pain reliever and a preventive medicine for heart attacks and strokes. It has come a long way from Edward Stone’s walk in the woods
59